Voxeleron at ARVO 2017

With ARVO 2017 upon us, we’re taking now the opportunity to reconnect with some of the many updates from the Voxeleron team. We hope that these are of interest and would be happy to hear back from you. Better still, we hope to see you in person in Baltimore! We’re at booth #3520, not far from ARVO central.
Orion™ Updates
Our OCT analysis software really has broadened as an offering with several updates and quite the road-map, as covered, amidst some news, in the following:
NIH SBIR Award
This February we were awarded a Phase I Small Business Innovation Research (SBIR) grant from the National Institutes of Health’s (NIH) National Center for the Advancement of Translational Sciences (NCATS), to further develop Orion as device-independent retinal image analysis software. We are obviously very proud to receive such a grant, and are already working hard toward achieving all of the specific aims outlined in our application.
Our main clinical collaborators are Dr. Pablo Villoslada (IDIBAPS/UCSF) and Dr. Pearse Keane (UCL/Moorfields), and we are very fortunate to have the support of such expertise. The grant will allow us to validate our retinal OCT image segmentation algorithms in cases of both ocular and neurological diseases, and also to add and validate change analysis functionality for longitudinal data. This work will be released into our Orion product by the Fall of 2017, further consolidating Orion’s place as the most advanced and only device-independent OCT analysis software commercially available. It is recognition of both the work that has been done and also the clear clinical need for such software.
“The Orion software was able to provide consistent and accurate segmentation of retinal layers and in addition a whole toolbox of added options to interpret and measure retinal parameters. We used the software extensively on patients with epiretinal membranes and macular holes, which are notoriously difficult for automatic analysis, with excellent results. This software was invaluable to our work as some features are simply unavailable anywhere else. We also would like to note the excellent support with prompt replies to all our questions.”
Introducing DeepSee™
The use of artificial intelligence (AI) in healthcare in general and in image analysis in particular is a very hot topic. And for good reason as, used correctly, deep learning-based algorithms are extremely powerful. In keeping with Voxeleron’s stated objective to be at technology’s leading edge, we will be demonstrating DeepSee, our recently developed deep learning architecture, in Orion at ARVO.
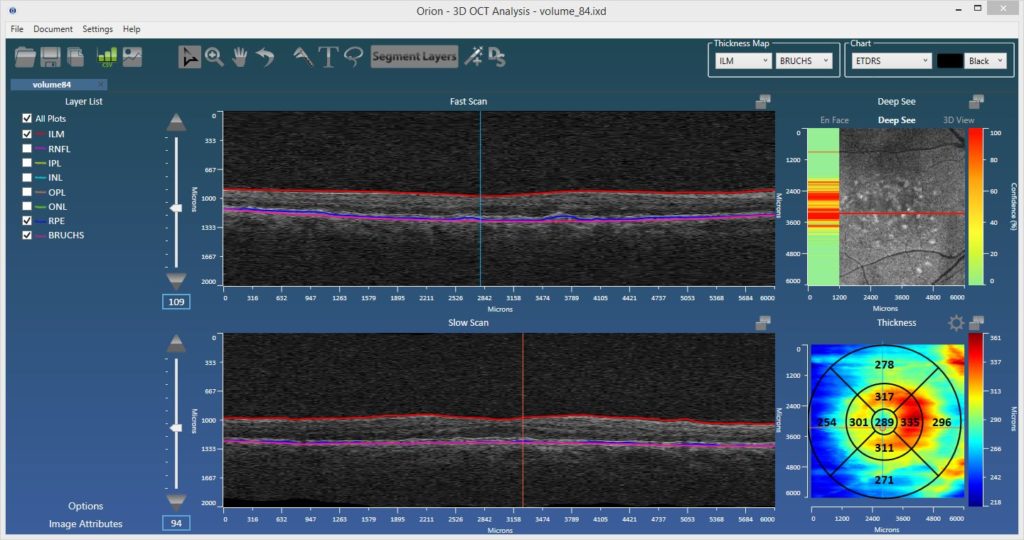
DeepSee uses deep, convolutional neural networks (CNNs) to classify the image data bounded by Voxeleron’s segmentation algorithms. As shown in our collaboration with Dr. Robert Chang of Stanford University (Poster B0415: “Deep convolutional neural networks for automated OCT pathology recognition”), this significantly improves accuracy compared to using the entire tomogram. We can then present to the user what data is suspected as being pathologic, facilitating faster image review and highlighting potential abnormalities that might not be seen otherwise. It’s also an orthogonal look at pathology that is not simply considering overall thicknesses.
These are powerful techniques that should complement and not define workflow, and this is our first step in this direction, as well as an ophthalmic industry first.
3dEdit™ – Intelligent Editing Wizard
This feature was first demonstrated at ARVO 2016, rolled out shortly afterward, and has received tremendous feedback. 3dEdit is patent-pending editing functionality that allows a user to manually edit or define entire surfaces in seconds. Correction of layers can be exceedingly laborious, so by improving editing efficiency it is possible to achieve huge savings in the time taken to perform these tasks. This can directly translate to significant cost savings for both reading centers and CROs.
Indeed, a study at Duke showed that the editing of the choroidal surface took on average 260 seconds per slice by trained evaluators! If you stop by our booth at ARVO, we’ll demonstrate exactly that task done in around 60 seconds per volume.
“The software has greatly sped up our research workflow, especially the new intelligent correction feature. The team at Voxeleron are great to work with and always ready to discuss potential future projects.”
Adam Dubis, Ph.D., Lecturer, Institute of Ophthalmology, University College, London (UCL).
3d Visualization
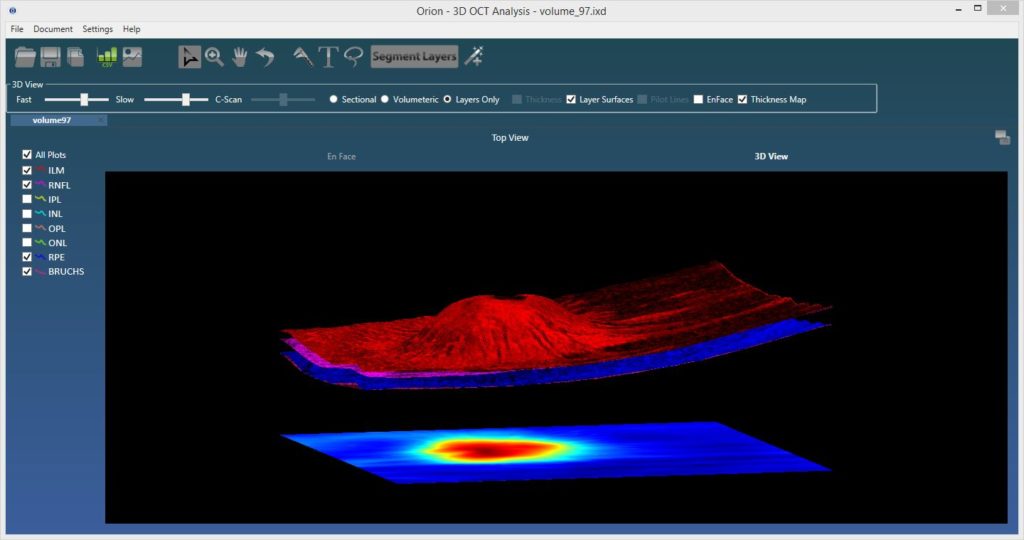
The ability to visualize OCT data in 3d was added toward the end of last year. The viewing supports rendering with and without segmented surfaces, cross-sectionally or by the full volume. It’s very cool, but perhaps more importantly, it lays the framework for some exciting new OCT angiography visualization approaches that we intend to release later in the year.
Angiography Support
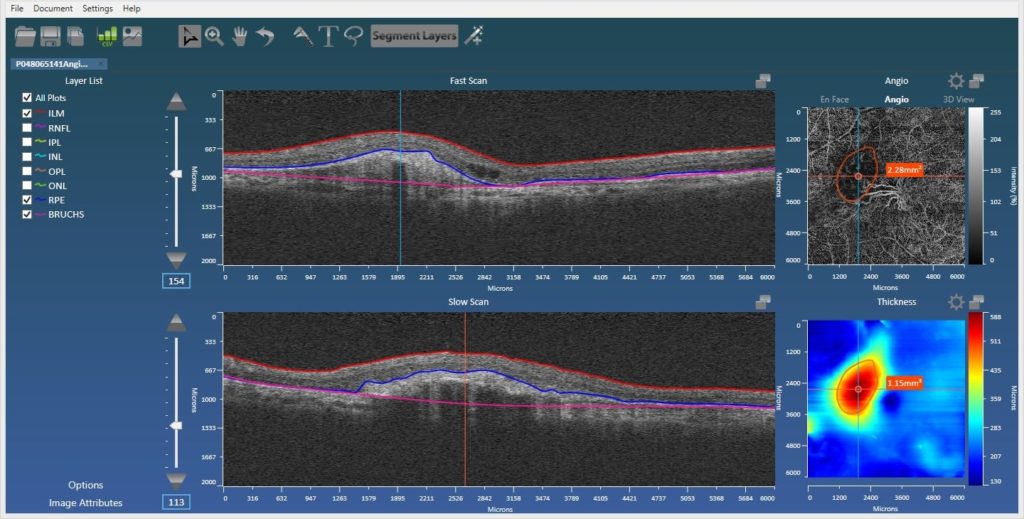
Also added is automatic support for the Zeiss AngioPlexTM device allowing the user to utilize Orion’s superior segmentation functionality to better view blood-flow within the relevant vascular beds. The user can customize these, but the standard superficial, deep, inner, and outer retinal plexuses are all there, as is the ability to view flow in the choriocapillaris, defined via our RPE-baseline segmentation. Projection artifact removal is also now integrated, so this new functionality, along with the ability to manually define regions of interest (ROIs – below), makes the Orion offering all the more compelling.
ROI analysis
In part to perform quantification in OCT angiography, but also to support volumetric and area measurements in structural OCT, we added free-form definition of ROIs in both en face views to report areas and volumes. This allows, for example, the user to delineate and measure regions of non-perfusion in angiography images, quantify the volume between disruptions to the RPE and the RPE-baseline, or define areas of atrophy as seen from the en face projection.
Orion in Press
We’re pleased to see some new journal publications featuring Orion that are now published:
- “Optical coherence tomography segmentation analysis in relapsing remitting versus progressive multiple sclerosis”, Raed Behbehani et al., PLoS ONE 12(2): e0172120. doi:10.1371/journal.pone.0172120.
- “The measurement repeatability using different partition methods of intraretinal tomographic thickness maps in healthy human subjects”, Tan J, Yang et al., Clinical Ophthalmology, Volume 2016:10 Pages 2403—2415, Nov 2016.
Orion Bits and Bobs
And here are some more updates that didn’t quite warrant their own heading:
- Throughput – the fastest just got faster! 8 layer segmentation on Cirrus 200-by-1024-by-200 cubes averages 2.2 seconds (i7-4900, and no GPU use) and just 3.3 seconds on the 512-by-1024-by-512 cubes! That processing speeds are linear with volumetric size bodes well for use with Swept-Source OCT devices that collect tons more data running at ~100 A-scans/second.
- Heidelberg Spectralis Integration – this is more complete and shows the fundus image aligned to the OCT data, along with synchronized navigation bars. It is very helpful to precisely see, for example, where features in the SLO image are relative to the OCT data.
- More formats – We already supported more image formats than any other application, yet we continue to add more devices. We are now pleased to say that we natively support all major OCT manufacturers!
- Change Analysis – as discussed in the piece on our SBIR grant, this is fully funded. Building on previously implemented methods, which include fully 3d OCT alignment, we look forward to incorporating feature packed change analysis tools, where any sector over any layer can be analyzed longitudinally in a practical and intuitive way.
- Roadmap – along with change analysis, more deep learning methods and an ever increasing list of supported data formats, we will also be working on visualization and quantification in OCT angiography. This is an exciting area, in much need of analysis tools, so do watch this space!
And Finally
While in Baltimore, we’ll be glad to visit the lab of Professor Joseph Mankowski, with whom we collaborate in automating methods of quantifying nerve fiber density and other metrics in corneal confocal microscopy. The original reference in this exciting collaboration is here, but the goal now is to take these methods clinical. Toward that aim, we also have some relevant updates (Poster A0383: “Automated analysis of in vivo confocal microscopy images of corneal nerves“), work done in conjunction with the Professor Mankowski’s lab and also Professor Charles McGhee’s lab at the University of Auckland.
If you cannot make ARVO this year do drop us a line instead. While we cannot guarantee we’ll save you the 2017 edition of the Voxeleron pen, we will be delighted to hear from you.
With best regards,
The Voxeleron team.

